| Bacteria
- Sensing the Environment |
Getting
about in the world
Bacteria
have a wide variety of means of locomotion, depending on the
type. Some bacteria are non-motile, relying entirely upon
passive flotation and Brownian motion for dispersal. However,
most are motile; at least during some stage of their lifecycle.
Motile bacteria move with "intent", gathering in regions which
are hot or cold, light or dark, or of favourable
chemical/nutrient content. This is obviously a useful attribute.
Many species glide across the substratum. This may involve
specific organelles, such as the filament bearing goblet-shaped
structures in the walls of Flexibacter (Moat, 1979). Often,
however, no specific organelles appear to be involved and
gliding may be attributable to the slime covering of some
bacteria or the streaming of outer membrane lipids of others
(e.g. Cytophaga (Moat, 1979)) or to other mechanisms currently
being elucidated. The spiral-shaped Spiroplasma corkscrews its
way through the medium by means of membrane-associated fibrils
resembling eukaryotic actin (Boyd, 1988). Gonogoocci exhibit
twitching motility (intermittent, jerky movements) due to the
presence of pili (fine filaments, 7 nm diameter, less than one
micrometer long) which branch and rejoin to form an irregular
surface lattice. However, more than half of motile bacteria use
one or more helical, whip-like appendages, about 24 nm diameter
and up to 10 mm long, called flagella (sing. flagellum).
Flagella would be of limited use if bacteria could not sense
their environment - they might help disperse cells to new
environments but energy would be better used if bacteria could
sense where they were going and respond appropriately. The
success of flagella is evident from their wide occurrence and
high expense (each flagellum comprises 1% of a bacterium's
protein (Neidhardt, 1987) and about 2% of its genome (some 50
genes) for their synthesis and control. Perhaps the main
advantage of flagella propulsion is their speed. Escherichia
coli is 2 mm long and has a single flagellum that propels it at
about 20 mm/s. In contrast speeds for gliding motility range
from 1-10 mm/s. Flagella, therefore, allow bacteria to respond
faster to changes in stimuli and enhance dispersal over a large
area. The pattern of flagellation varies with each species. Some
classification schemes divide flagellated bacteria into two
groups: the Pseudomonadales have one or more flagella at one or
both poles of the cell (they are polarly flagellated). The
Eubacteriales have a random distribution of flagella, sometimes
covering the whole surface of the cell (they are peritrichously
flagellated).
Chemosensing
mode in Escherichia coli
Bacteria
can move towards or away from stimuli. In the vast majority of
bacteria (all except the very largest) this occurs by
chemokinesis, when the stimulus is a chemical such as a food
substance or a noxious irritant. Chemokinesis is an indirect
process in which the bacterium alternately swims for a stretch
and then changes direction and changes its frequency of turning
in response to environmental stimuli, which it can be
demonstrated by computation leads to the bacteria gathering near
food sources whilst avoiding noxious stimuli. Note that the
definition used here of chemokinesis is in keeping with
classical zoological and botanical definitions, but
bacteriologists usually define the terms differently and refer
to the phenomenon just described as chemotaxis. Call it what you
like, the mechanism will now be described.
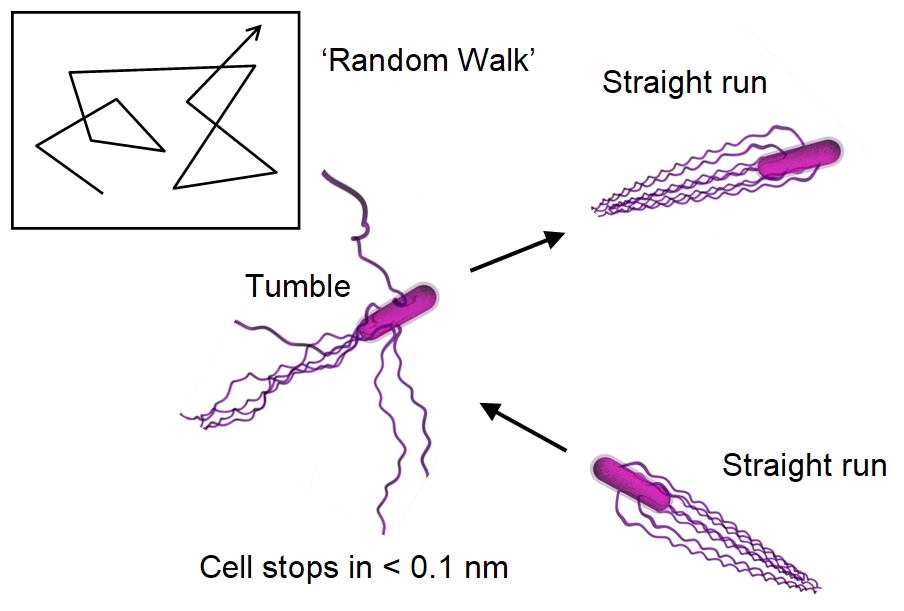
When unstimulated, free-swimming Escherichia
coli
swarmer cells make long straight runs with infrequent tumbles
(see figure below). During
the straight runs, the flagella rotate CCW (counter-clockwise) and the flagella come
together and work as a single bundle, possibly driven together
by hydrodynamic forces (note that the cell body rotates in the
opposite sense, CW, to the flagella). During
the tumbles, the flagella reverse rotation to the CW
(clockwise) sense,
the wavelength of their filaments also changes and the pitch
increases and the flagella mechanically uncouple and fly apart
and the cell tumbles (rotates randomly). Chances are that after
a tumble when the flagella resume CCW rotation and reform a
bundle that the bacterium will start off in a new direction: it
has turned by a random number of degrees. Recent evidence shows
that only one or two flagella in a bundle are required to
uncouple to effect a tumble.
Below:
Patterns of movement in Escherichia coli
What do we mean by frequency of tumbling? The bacterial flagella have a switch mechanism, alternating between two states: CW rotation and CCW rotation, with the rotation rate remaining more or less constant. The Gibbs free energy for the CCW (swimming) to CW (tumbling) transition is about +40 kJ mol-1, and this energy is provided for by random thermal fluctuations. When the cell is triggered to tumble more, it simply increases the probability of tumbling by reducing the free energy required, making it more likely that thermal noise will provide the necessary energy.

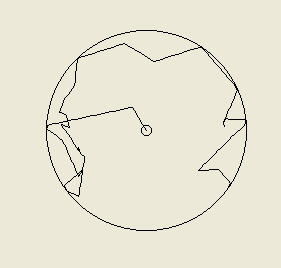
Above left - a computer-simulated 'random walk' of a bacterial cell starting at the center of the arena and tumbling at random intervals. Right - two such simulated cells tracked over 15000 seconds each. Despite starting at the center they soon reach the edge of the arena (by random chance); they have been programmed to bounce of the walls at the arena's edge.
When Escherichia coli nears the source of an attractant chemical, moving up a concentration gradient, it's tumbling frequency decreases and it makes longer runs. If tumbling frequency was simply inversely proportional to attractant concentration, then computer simulations reveal that this will actually lead to a repellent effect: the bacterium will tend to move away from the attractant source. However, when the computerized virtual cell is given a memory, allowing it to compare the concentration at different points in time, it is able to measure the gradient rather than simply the concentration. By reducing tumbling as it moves up the gradient toward the attractant source, then it will indeed become 'attracted' to the source (meaning that it spends longer near the source). This is indeed found to be the case: bacteria have a memory of a few minutes that allows them to make temporal comparisons of chemical concentrations in their environment.

Computer-simulated bacterial cells programmed to respond to the
(exponential) concentration gradient of a chemo-attractant placed in
the center of the arena. Each cell has been given a one second
memory with which to measure changes in attractant concentration
over time. The cells tumble less if they move up the concentration
gradient and tumble more if they travel down the gradient (within
preset minimum and maximum tumbling frequencies). Each picture shows
4 cells traced for 5000 seconds each.
Left - each cell started at the edge of the arena at the bottom
right corner at about 5 o'clock. Right - the bacteria start nearer
to the arena center. Note how the bacteria remain near the
attractant even though they are in constant motion.
Repellents work in a converse manner - the bacteria tumble more frequently as they approach the repellent (as the gradient increases) and so tend to avoid it:

Computer-simulated bacteria programmed to respond to a chemo- repellent at the center of the arena, by tumbling more frequently if they travel up the gradient and tumbling less frequently if they travel down the gradient (within preset minimum and maximum tumbling frequencies). Each cell had a one second memory to enable temporal comparisons in repellent concentration. Left - one cell starting at the center and was tracked for 15000 seconds. Right - 3 cells began in the center of the arena and were tracked for 15000 seconds each. They can be seen to clearly avoid the repellent and stay near the edge of the arena.
Chemosensing
modes in other flagellated bacteria
Some
alpha-subgroup bacteria also have a CW/CCW flagella switch, for
example in Caulobacter
crescentus
CW rotation leads to swimming, CCW rotation to tumbling (the
opposite sense as in Escherichia
coli).
Others, however, have flagella that rotate in one direction
only. In other members of this group very different strategies
are employed. Rhodobacter
sphaeroides has
a single right-handed flagellum in a lateral position which can
push the cell at up to ~35 mm/s. This flagellum rotates only CW
(or CCW in some variants). If an anaerobically grown cell
detects a decreasing gradient of chemoattractant (if it is
moving from the source of the attractant) then its probability
of turning increases (note that this response is opposite in
sense to that of E. coli, but equivalent). During
turning, the flagellum stops and assumes the form of a planar
spiral. This whole flagellum spiral slowly rotates, turning the
cell by a random amount. Sinorhizobium
meliloti
has 2-6 short, peritrichously or lophotrichously arranged
right-handed flagella with fixed CW rotation, during which they
form a propulsive bundle. The flagella rotate continuously,
however, asynchronous rates of rotation cause the bundle to fly
apart and the cell to tumble. more information on these other
sensory response modes can be found here.
Mechanism
of chemosensing
This
section of the article is somewhat more technical.
The best-studied chemosensing mechanism of Escherichia
coli
is the Che
phosphorelay system
summarised in the diagram below. In Escherichia
coli
the chemosensors are methyl-accepting chemotaxis
proteins (MCPs)
that span the inner membrane as dimers (pairs). These are
connected to a phosphorelay system. In such phosphorelay
systems, the signal is relayed by protein
phosphorylation,
which involves enzymes called phosphatases (which remove
phosphate) phosphorylases (which attach
phosphate) and kinases (which transfer
phosphate from one molecule to another). These receptors tend to
be concentrated at the cell poles. Together with several of the
chemotaxis (Che) proteins they form a MCP:CheW:CheA:CheY
receptor complex.
When
stimulated by a repellent
CheW functions as a relay
linker, connecting the MCP to CheA. When an MCP is stimulated by
a chemorepellent binding to it, the receptors cluster in groups
of about 7 MCP dimmers together with about 7 CheA dimers and are
in their activated state. This state activates CheA which
autophosphorylates (phosphorylates itself) to CheA-P.
CheA-P is an active kinase
and phosphorylates CheY to the active CheY-P form.
CheY-P then dissociates from
the receptor complex and diffuses to the flagella motor where it
binds FliM in the C ring (see flagella structure) and it also
binds the regulator phosphatase CheZ which undergoes
delayed activation as it oligomerizes to its active form. This
binding to CheZ increases
the probability of tumbling (CW rotation)
by reducing the activation energy of the switch. After a delay
CheZ becomes active and this acts as a phosphatase, removing
phosphate from CheY-P to yield CheY which reassociates with the
receptor complex. Thus, CheZ acts as a time-delay which resets
the system, a form of adaptation which helps confer the memory
needed for the system to respond to concentration gradients.
A second adaptation system also exists, which is
methyl-dependent. CheA-P not only phosphorylates and activates
CheY, but it also phosphorylates and activates CheB, at a slower rate,
producing CheB-P after a time-delay. CheB-P is a specific
methyl-esterase which cleaves a methyl group from MCP (MCP-Me →
MCP). The methylated form is the active form which stimulates CW
flagella rotation and tumbling. The demethylated form becomes
inactive and CCW
rotation and straight runs become favoured again - returning the cell
to its pre-stimulus state. Chemorepellents thus stimulate MCP
demethylation. This periodic resetting of the receptor occurs
every few minutes, giving the cell a memory of an equivalent
duration. CheR also helps to reset the system by antagonising
CheB-P as CheR is a methyl transferase which methylates MCPs.
Above: Che phosphorelay signalling system. Binding of a chemical repellent (ligand) to the receptor (MCP) increases tumbling frequency when repellent concentration increases. The protein CheW (W) connects the MCP to the protein CheA (A). When the repellent binds MCP, CheA becomes activated by autophosphorylation and then phosphorylates CheY, activating it too. The active phosphorylated cheY diffuses to the flagellar motor and binds to it, along with CheZ to which the active CheY also binds. This increases the probability of tumbling by tending to switch the flagella into CW rotation mode.
When
stimulated by an attractant
Attractants
work in a converse manner. When an attractant binds to the MCP,
the receptors deactivate and de-cluster into separate dimmers
(with associated CheA dimers). This inhibits the
autophosphorylation of CheA, deactivating it and reducing the
concentration of CheY-P, resulting in an increased probability
of CW flagella rotation and a reduced probability of tumbling.
Sugars are chemoattractants that do not bind MCPs directly.
Instead they either bind their sugarspecific periplasmic
binding protein (PBP)
which is involved in sugar transport into the cell as well as
chemotaxis. [Recall that the periplasm is the region between the
inner and outer membranes in the envelopes of gram negative
bacteria and contains various PBPs which bind to nutrients.]
Galactose, maltose and ribose work this way. At least the PBP
for maltose is known to be localised at the cell poles along
with the MCPs. Alternatively, some sugars, e.g. glucose, mannose
and mannitol, bind to a specific Enzyme II of the PEP-dependent
sugar phosphotransferase system (PTS) (PEP is
phosphoenolpyruvate). The PTS Enzyme I modulates the kinase
activity of the MCP-CheW-CheA receptor complex by
phosphorylating CheA and simultaneously phosphorylates the sugar
as it is imported through the EII/EIII proteins. [More
explanation to be added here.]
MCPs also respond to changes in pH and temperature. Oxygen acts
as an attractant for Escherichia
coli
and its receptor is Aer (an MCP homologue) which is cytoplasmic
and lacks methylation sites for adaptation. Aer also binds FAD
(flavine adenine dinucleotide, an electron/hydrogen carrier used
in respiration) which acts as an internal signal for the redox
state of the electron transport
chain
(ETC).
A summary of this complex process of chemosensing in enteric
bacteria, such as Escherichia
coli,
is shown in the figures and table below:
Attractants bind either directly to MCPs or via periplasmic binding proteins, the latter serving as adapters to increase the range of chemical ligands to which the MCPs can respond. The chemosensory ligand must first diffuse across porins (protein pores or channels) in the outer membrane (OM). IM: inner membrane; OM: outer membrane; PP: periplasm.
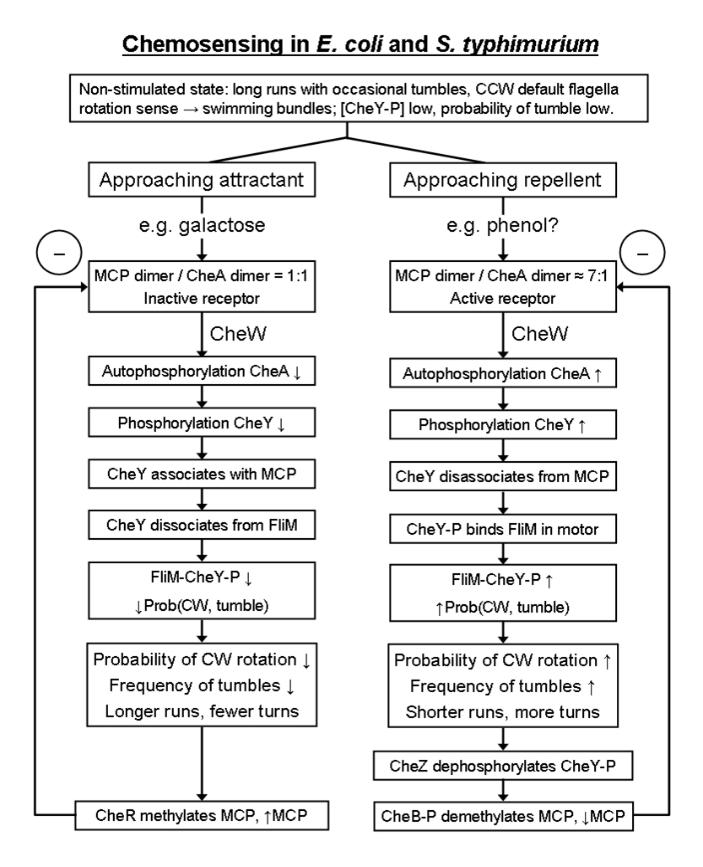
Above: when a repellent (or PBP + attractant) binds the Escherichia coli MCP, such as phenol, or equivalently when an attractant dissociates from the MCP, then CheA autophosphorylates itself and then passes the phosphate on to a second messenger called CheY (CheA may potentially phosphorylate several CheY molecules in this way). Phosphorylated CheY, CheY-P, is active. CheB also becomes activated by phosphorylation. This is the characteristic two-component system (TCS) using a phosphorelay mechanism, which is characteristic of signalling in bacteria (compare this to the eukaryotic kinase cascade). CheW is an adaptor protein, attaching CheA to the MCP so CheA can respond to ligand binding the MCP. CheB-P demethylates the MCP. The more methylated the MCP, the more it activates CheA. Thus, demethylation by CheB (an esterase) reduces activity in the system by reducing CheA phosphorylation. This resets the MCP sensor should the concentration of repellent remain unchanged, this is adaptation to the stimulus and is a form of memory. It would be interesting to see to what extent adaptation occurs to repellents compared to attractants. (There are few repellents in the Escherichia coli case and establishing their existence has been problematic, but other bacteria may possibly sense a wider range of repellents). This adaptation is brought about by MCP methylation / demethylation which depends on the enzymes CheB and CheR.
Above:
when an attractant (or PBP + attractant), such as a specific
sugar like galactose, binds the Escherichia
coli
MCP, CheA phosphorylation and activation are reduced and so
phosphorylation and activation of CheY is also reduced.
Similarly phosphorylation and activation of CheB is reduced.
CheR (a methyl
transferase)
remains active at a more-or-less constant rate regardless and is
now able to demethylate the MCP without competing with CheB.
Methylation of the MCP is thus reduced, which tends to increase
the activity of the MCP and CheA, resetting the system should
the concentration of attractant not change. This is adaptation,
which is crucial if the cell is to respond to changes
in stimulus concentration.
Note: much of the details
of this system are still being elucidated by experiment. There
is some uncertainty as to the mode of CheW and CheA binding -
CheW is described as linking the MCP to CheA by some sources,
whilst others claim that CheA and CheW bind to MCP
competitively. Here we present only models (the so-called 2:2:2
model) which will be altered as and when my own researches turn
up positive developments. It has also been suggested that CheA
binds between the two MCP dimers, facilitating their
dimerisation.
Effect
of CheY
When
CheY is phosphorylated to CheY-P by CheA, it becomes active. In
this active form it disassociates
from the MCP receptor and binds to FliM, a protein component of
the C ring in the cytoplasm (see bacterial
motility for a description of
the flagella structure). This switches the rotation bias of the
flagellum to CW
rotation, which favours tumbles. CheY-P is phosphorylated when
the concentration of a repellent increases, or equivalently,
when the concentration of an attractant reduces.
Notes: CheR is active at the same basic rate in both cases (that is it is not affected by binding or detachment of attractant or repellent from the MCP). This will methylate the MCP at a basal rate. However, when the concentration of repellent increases (or concentration of attractant decreases) then CheB is phosphorylated to CheB-P and activated. CheB-P demethylates the MCP, opposing the action of CheR. Thus, when active, the methylation of MCP decreases. Binding of repellent initially activates the MCP, then the delayed action of CheB-P deactivates it, leading to adaptation of the response.
Color and label your own diagram of Escherichia coli and its nanobrain! (Note these diagrams are not to scale: the size of the signaling components have been exaggerated for clarity).
For more on chemokinesis in bacteria:
Integral feedback control and asymptotic tracking
One key feature of this signaling pathway is the aforementioned property of adaptation. The E. coli chemotactic system is dynamic: it responds primarily to changes in ligand (attractant or repellent) concentration rather than the absolute concentration of ligand. This is important since the bacterium wants to move closer to an attractant, such as a food source, and further away from a source of repellent: it is the concentration gradient that must be sensed as already discussed. The key thing about adaptation is that after a stimulus, such as an increase in repellent concentration activates the MCP receptors then after a certain short time interval the receptors return to their preset (resting) level of activity even if the ligand concentration remains elevated. Similar arguments apply to a decrease in attractant concentration activating the system or an increase in attractant / decrease in repellent inhibiting the system: the system detects the change then returns to its set level. This means that if repellent concentration increases from say 2 (in arbitrary units) to 4, and remains at 4, the system responds initially to the change from 2 to 4 and then returns to its preset level of activity.
A similar adaptation (though by different mechanisms) occurs with the human olfactory system. If you enter a smelly room, perhaps one in which the rubbish needs taking out, then you will be greeted by the unpleasant odor. If you remain in the room then after a while you are no longer aware of the odor: your olfactory system has adapted and returned to its preset level. If you go outside for a bit, into fresh air, and then return, however, then you will notice the smell again, for a while. Your olfactory system is specialized to detect changes in odor profiles rather than background odors: the former indicate change and changes are generally much more important. For a more sensitive animal, a new odor or an increase in the odor might indicate an approaching predator!
Adaptation returns the system to its preset level. This preset level is that at which the system is most sensitive to further changes: adaptation not only filters out unimportant constant background signals, but also ensures that the sensitivity of the system to further changes is maintained. In other words: the output from the sensory system is independent of the input level (stimulus of ligand or odor) in steady-state (when environmental signals are constant). Biologists call this perfect adaptation, engineers call it asymptotic tracking.
The most intuitive way to compensate for an error is to generate an output signal proportional to the size of the error, what we call a proportional controller. As an example, consider a heater attempting to keep the temperature of a room at the set temperature. If the temperature drops too low the heater comes on and switches off once the error = set temperature - actual temperature. However, the elements will remain hot for a while and so overcompensate. At best the actual temperature will oscillate around the set temperature. The residual or steady-state error that causes such overshoots or oscillations can be prevented by using integral control, which is employed in most actual heaters. Integral control is necessary to achieve asymptotic tracking, in which the system returns precisely to the set-level. An integral is essentially a sum of many small changes (not an exact definition) and integral controllers generate an output based on how the error changes over a period of time: they respond to the gradient or rate of change of the input signal. This is precisely what E. coli does: it samples ligand concentration over an interval of time (and space) To determine the gradient of the change and it is this gradient the bacterium responds to. Precise adaptation or asymptotic tracking is brought about by an integral controller. A third control system commonly employed in engineering is the differential controller, which uses collected input data to predict future trends: is the temperature going up or down, for example. Some control circuits may combine all three methods as a PID controller.
To achieve asymptotic tracking, it has been shown that the CheB / CheR system must have the following properties: the rate of receptor methylation must depend on receptor activity and not methylation level and not directly on ligand concentration; only inactive receptors should become methylated and only activated receptors demethylated. Indeed, this appears to be the case in E. coli, at least to a first approximation (Yi et al., 2000). Once again, Mother Nature has already beaten engineers to it! Nevertheless, the fact that both devise similar systems is reassuring.
For more details on adaptation see the pdf on network analysis of the Escherichia coli chemotactic system.
Geometry of the sensory array and cooperativity
Early models used to think of signaling pathways, such as the MCP-CheY pathway in E. coli, as consisting of molecules simply diffusing around at random, either in the volume of the cytosol of the cell or in a cellular membrane, and reacting when they encountered one-another. Although diffusion still plays a role, of course, more and more it is being realised that biomolecules form multimolecular complexes or nanomachines. Large molecules in particular, such as proteins often associate with one-another in such a way that one protein is not too far away from other proteins it must react with. First of all, this changes the kinetics: it may force proteins to react in certain stoichiometries (ratios) rather than reacting with a large number of diffusing individual proteins. For example, in eukaryotic cells proteins that make up a signaling module called a kinase cascade would ordinarily form an amplifier when the components are free to diffuse, but if the components are held into a multiprotein complex by adaptor or scaffold proteins the module acts as a noise filter instead. (A scaffold protein can be thought of as a large multi-adaptor protein that can bind to several other proteins and hold them in place). In bacteria, with their small cell size and simpler architecture, we might expect such complexes to occur less often, but they do occur and in E. coli and some other bacteria that have been studied, the chemotactic receptors form multiprotein complexes.
Above:a model of the chemosensory array of E. coli as seen when looking down onto the membrane (into the cell). The receptors are shown in green as trimers of dimers (pointing out of the page); the CheW adaptor proteins are shown as small blue circles: each dimer is strongly associated with one CheW; orange circles are CheA dimers. The diameter of the central 'pore' is 10 nm (1 nm = 10-9 m). The stoichiometry is 6MCP:3CheW:3CheA. Many of these hexagonal arrays are fitted together in a 2D sheet containing something like 1500 to 4500 MCP receptors (up to 125 of these hexagonal units) - the actual number is probably dynamic. This would form a patch about 0.2 to 0.6 μm in diameter (1 μm = 10-6 m). This field of sensors occurs at one or both ends in rod-shaped bacteria like E. coli. The large MCP proteins cause the membrane to curve and so to minimize energy this arrays assemble at the cell poles where the curvature is greatest. (Based on: Shimizu et al., 2000).
Below: an alternative representation:
Such complexes of receptors raises another possibility: cooperativity. Cooperativity between the receptors occurs when a ligand binding to one receptor increases the affinity of the other receptors and so enhances the binding of additional ligands. (Cooperativity may also involve the CheA, in which activation of one CheA increases the likelihood that other CheA units become activated when further ligands bind the receptors). The chief effect of this kind of cooperativity is to filter out noise from the signal. Noise can be external, e.g. the binding of a stray ligand, or internal, e.g. a CheA molecule momentarily flipping into its active state with low probability in the absence of ligand (thinking in terms of repellents to make things easier). This arrangement may also possibly amplify the signal, but note there is a trade-off between 'sensitivity' and noise reduction. The chemosensory apparatus of E. coli certainly filters out noise efficiently and also amplifies the signal considerably and these protein complexes are almost certainly integral to how this is done, although the exact mechanisms are not yet fully understood.
For more details of these phenomena see our pdf on modeling this system computationally using network analysis.
Responses
to other stimuli
Bacteria
can also respond to a variety of other stimuli, apart from
specific chemicals, and responses to temperature, light,
osmolarity, pH and touch have been observed. Photosynthetic
bacteria may move towards the light to photosynthesise, or away
from it if the light is too bright and potentially damaging. One
particular response we shall look at here is the response to
magnetism, in particular to the Earth's magnetic field. Bacteria
that move magnetic North or South in a magnetic field are called
magnetotactic.

Above: Magnetospirillum (Aquaspirillum)
Magnetotactic
bacteria
are Gram-negative, flagellated and motile bacteria. They swim at
about 100 mms-1. They are microaerophilic, aquatic
and contain magnetosomes. These bacteria come from many
taxonomic groups. The magnetosomes are intracellular magnetic
grains that give each cell a permanent magnetic dipole moment.
The local geomagnetic field (about 0.5 G) places a magnetic
torque upon the bacteria, aligning them with the magnetic field.
Magnetospirillum
magnetotacticum
(Aquaspirillum magnetotacticum) has Fe3O4
magnetite (lodestone) magnetosomes. Each cell contains about 20
cuboidal-octahedral crystals. Each crystal is about 420
angstroms (42.0 nm) in diameter and the crystals are arranged in
a chain. Single magnetic domains in magnetite range from
400-1000 angstroms. Full alignment requires the ratio of the
interactive magnetic energy with the applied field to the
thermal energy to be greater than about 10. A ratio of 16 is
obtained for 22 500 angstrom particles. As we shall see, there
are other mechanisms that stabilize the structure and keep the
crystals aligned.
Thus, the magnetosomes function as a ferromagnetic biocompass. At 0.5G and 30oC,
80-90% of the cells are fully aligned. The cells clump together
if they contain too many magnetosomes. Recorded levels of
magnetotactic bacteria yield 103-104 cells
/ ml of slurry in aquatic environments (in New England). These
bacteria are absent from heavily polluted waters and absent in
limestone caverns, thermal springs, thawed Antarctic sediments
and iron-rich seeps. They are abundant in water purification
plant settling basins, ponds with organic sediments, and
sewage-treatment oxidation ponds.
In the Northern Hemisphere, magnetotactic bacteria swim
predominantly northward, whilst in the Southern Hemisphere they
swim predominantly southward. The vertical component of the
geomagnetic field, due to Earth's curvature, is directed upwards
in the Southern Hemisphere, is zero at the geomagnetic equator,
and is directed downwards in the Northern Hemisphere. Therefore,
the bacteria are directed downwards in both hemispheres and so
are abundant in sediments, but absent in surface water. At the
geomagnetic equator there are approximately equal proportions of
each polarity and the bacteria swim horizontally.
Magnetospirillum
magnetotacticum is a microaerophilic
chemoheterotroph that metabolises organic acids, like fumaric,
tartaric and succinic acids as its sole carbon (and energy?)
source. Nitrate is the principle electron-acceptor for
microaerobic growth on tartrate. Traces of oxygen are required.
(As a substrate for oxygenases? For haem production?). Nitrate
is reduced to nitrous oxide and ammonia in the process. Thus,
this bacterium is a denitrifier. In low oxygen
tensions (< 0.2 kPa) these bacteria also reduce ethyne. In
high oxygen tension (above about 6 kPa) they do not produce
magnetosomes and are not magnetotactic. This bacterium lacks
catalase activity and so is sensitive to high oxygen
concentrations. They have polar or bipolar flagellation. Pili on
coccoid forms, extracellular polysaccharide on spirilla, allow
them to adhere to sediment particles.
Magnetosomes are typically (though not always) arranged in one
or more linear chains. Each magnetosome particle is a single
crystal of magnetite or greigite enclosed in a phospholipid
bilayer membrane. This membrane is derived by invagination of
the inner membrane but contains specialized proteins. This
causes some confusion in nomenclature - does 'magnetosome' refer
to the whole chain or just to one of the particles? Each
magnetosome particle/vesicle is a tiny magnet and the particles
within each chain are aligned -(N-S)-(N-S)-(N-S)- which helps to
stabilize the chain as a magnetic north pole is attracted to a
magnetic south pole. However, as anyone who has ever played with
magnets will know, this arrangement is metastable, meaning that
a slight displacement of one of the magnets can cause the whole
chain to collapse into an aggregated cluster of magnets.
Additional structures are needed to stabilize this arrangement
(see the diagram below). In Magnetospirillum the actin homologue
protein MamK forms a protein filament track along which the
magnetosome particles are aligned (see below). There is evidence
to suggest that the protein MamJ forms linkers that connect each
vesicle to the track. It is also possible that additional
structures link neighbouring magnetosome vesicles together. (See
review by Thanbichler and Shapiro, 2008).
Above: The magnetosome and associated stabilizing structures in Magnetospirillum.
Some
bacteria have two chains of magnetosomes at opposite sides of
the cell and it is possible that these are positioned by their
mutual magnetic repulsion. Large cells require a larger torque
to turn them and the large rod- shaped cells of Magnetotacticum
bavaricum,
which are 8-10 mm long, have 200-1000 crystals arranged in 5
chains. However, it is possible that this produces far more
torque than needed and magnetosomes may have additional
functions.
The individual crystals of magnetite are 35-120 nm in diameter.
A magnetite crystal contains one or more domains - regions in
which the molecules all have their magnetic moments aligned
parallel, reinforcing one another. If the crystals were much
larger than domain boundaries would occur, where the alignment
of the molecules switches between crystal domains with magnetic
poles pointing in different directions (typically opposite one
another). A crystal with several domains whose magnetic moments
tend to oppose and cancel one another will have a reduced
magnetic moment. If the crystals were much smaller, then they
would lose their permanent magnetism, due to the thermal motion
of the atoms. Thus, the size chosen is optimal for maximizing
the intensity of the crystal's magnetic field.
Some bacteria use greigite (Fe3S4) instead
of magnetite. In particular these bacteria occur in environments
where oxygen availability is low and sulfur-availability high.
Greigite is less effective, however, as it makes a weaker magnet
(one third as strong). At least one known species of bacterium
has crystals of both greigite and magnetite (a given magnetosome
particle must be one or the other and mixed crystals do not
occur). Such bacteria encounter environments of variable oxygen
content - if oxygen is scarce then they incorporate sulfur
instead.
Phototaxis
Photosynthetic
bacteria require light and are thus sensitive to it and, if
motile, able to locate the best region of illumination to
maximize their photosynthesis. For example the cyanobacterium Synechocystis
will crawl towards light by extending type 4 pili (T4P
or Tfp) which adhere to the substrate and then retract, dragging
the cell forwards. Evidence suggests that the whole spherical
cell acts as a lens to focus the light in order to determine the
direction towards the light source, as illustrated below:
This begs the question: how small can a lens be and still focus light? Microlenses have been manufactured down to about 10 μm diameter. As a first 'rule of thumb' one might expect a lens to have to be at least ten times the wavelength of light in diameter to work as a conventional lens. For blue light at 500 nm wavelength, this would require a lens of 5 μm diameter or greater. A typical cell of Synechocystis, however, is only about 2 μm in diameter. This does not mean that it can not refract and focus light, only that diffraction effects will become very significant and perhaps a sharp image could not be formed. However, it only needs to focus the light partially to create a measurable difference in light intensity around the circumference of the cell (assuming the photoreceptor to be in or adjacent to the cell envelope). This prokaryote is probably about as small as it can be and still perform this function effectively. The direction opposite to the spot where the focused light is brightest is the direction the cell needs to move in (towards the right in the above diagram).
Related Articles:
Network analysis of
the E. coli chemotaxis signaling pathway (pdf)
Evolution
of Bacterial Flagella
See also quorum sensing - how bacteria sense the presence of one-another!
Article updated 16/9/14, 12/7/2020, 16/7/2023
References:
Shimuzu, T. S., Le Novėre, N., Levin, M.D., Beavil, A.J., Sutton, B.J. and Bray, D. 2000. Molecular model of a lattice of signalling proteins involved in bacterial chemotaxis. Nat. Cell Bio. 2: 792-796.
Yi, T.-M., Huang, Y., Simon, M.I. and Doyle, J. 2000. Robust perfect adaptation in bacterial chemotaxis through integral feedback control. PNAS 25;97(9):4649-53. PMID:10781070 PMCID:PMC18287 DOI:10.1073/pnas.97.9.4649