| Bacterial Behaviour - Chemokinesis / Chemotaxis |
Examples - patterns of flagellation and patterns of movement
The following examples are illustrated in the figure below. The bacterium Chromatium okenii has about
40 flagella at one pole. It swims predominantly with the flagella trailing behind, but apparently at random,
the cell backs up, moving bundle-first several body lengths, then swims forward once more.
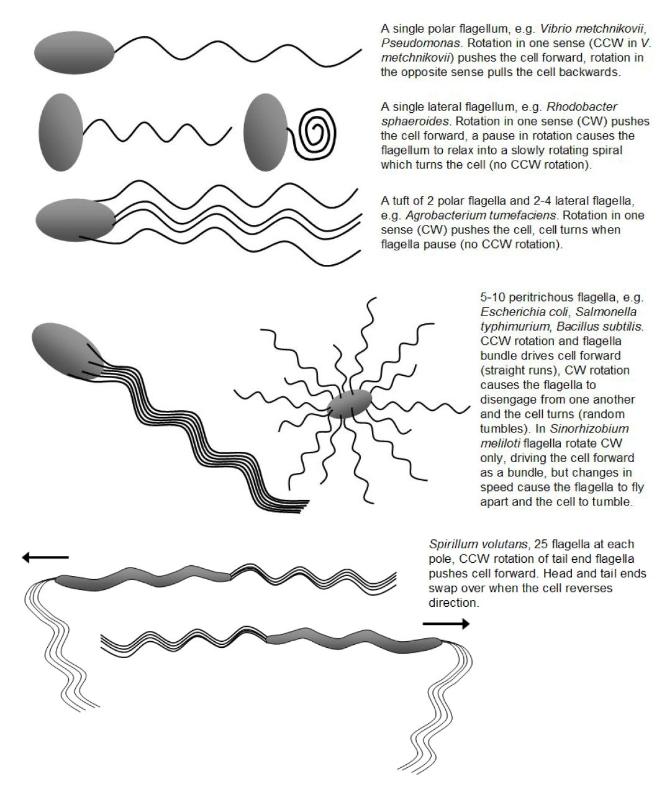
Cells with a single polar flagella, e.g. Vibrio metchnikovii, simply back up, and can swim equally well in
both directions.
Spirillum volutans has about 25 flagella at each pole and flips its bundles from a head-tail configuration
to a tail-head configuration and swims off in the opposite direction.
Escherichia coli is peritrichously flagellated and swims forward by bringing its six flagella together into a
flagella bundle. At intervals it stops moving as its flagella fly out in all directions and the cell tumbles,
thereby bringing about a random change in direction after which they resume smooth swimming. These
changes are brought about by reversals in the direction of flagella rotation - when the flagella rotate
clockwise (CW) their filaments come together into a single bundle at one end (the tail end) of the cell
(made possible because of the flexible hooks - the points of attachment of the flagella to the cell do not
change) and propel the cell in a smooth forward direction, but when the flagella rotate counterclockwise
(CCW) the bundle flies apart and the cell tumbles. The likelihood of a change in direction is biased by
sensory perception. As bacteria swim up a spatial gradient of an attractant, such as the nutrient
aspartate (an amino acid) the probability of reversal or change in direction reduces. When bacteria swim
away from an attractant, the probability of a change in direction increases to its normal level (for an
isotropic solution). Thus the bacteria eventually swim up the gradient because the runs are longer in the
favourable direction. Note that the cells must have sensors to detect (smell or taste) the nutrients in the
water in order to determine where they are coming from.
The following examples are illustrated in the figure below. The bacterium Chromatium okenii has about
40 flagella at one pole. It swims predominantly with the flagella trailing behind, but apparently at random,
the cell backs up, moving bundle-first several body lengths, then swims forward once more.
Cells with a single polar flagella, e.g. Vibrio metchnikovii, simply back up, and can swim equally well in
both directions.
Spirillum volutans has about 25 flagella at each pole and flips its bundles from a head-tail configuration
to a tail-head configuration and swims off in the opposite direction.
Escherichia coli is peritrichously flagellated and swims forward by bringing its six flagella together into a
flagella bundle. At intervals it stops moving as its flagella fly out in all directions and the cell tumbles,
thereby bringing about a random change in direction after which they resume smooth swimming. These
changes are brought about by reversals in the direction of flagella rotation - when the flagella rotate
clockwise (CW) their filaments come together into a single bundle at one end (the tail end) of the cell
(made possible because of the flexible hooks - the points of attachment of the flagella to the cell do not
change) and propel the cell in a smooth forward direction, but when the flagella rotate counterclockwise
(CCW) the bundle flies apart and the cell tumbles. The likelihood of a change in direction is biased by
sensory perception. As bacteria swim up a spatial gradient of an attractant, such as the nutrient
aspartate (an amino acid) the probability of reversal or change in direction reduces. When bacteria swim
away from an attractant, the probability of a change in direction increases to its normal level (for an
isotropic solution). Thus the bacteria eventually swim up the gradient because the runs are longer in the
favourable direction. Note that the cells must have sensors to detect (smell or taste) the nutrients in the
water in order to determine where they are coming from.
What is a spatial gradient of an attractant?
Escherichia coli is attracted to the nutrients it needs, such as the amino acid aspartate. This amino acid
could be coming from a lump of decaying organic matter in your large intestine, where Escherichia coli
lives. These bacteria will then gather around the food source to feed (we will look at how bacteria ingest
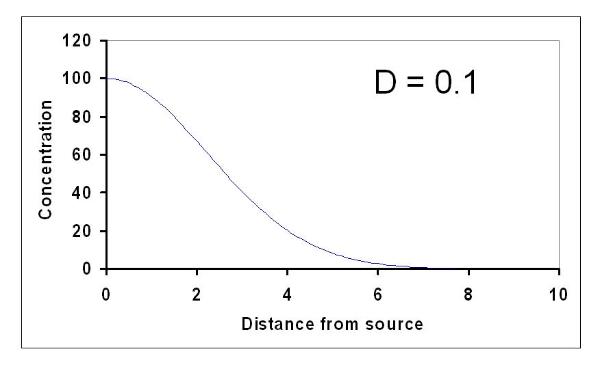
nutrients later). The nutrients will slowly diffuse away from the source into the surrounding water
(intestinal fluid) in such a way as to produce a diffusion gradient (or slope in concentration) with the
concentration increasing closer to the source and eventually dwindling to zero far from the source. Such
gradients are generally what we call exponential gradients, like the ones shown below:
Escherichia coli is attracted to the nutrients it needs, such as the amino acid aspartate. This amino acid
could be coming from a lump of decaying organic matter in your large intestine, where Escherichia coli
lives. These bacteria will then gather around the food source to feed (we will look at how bacteria ingest
nutrients later). The nutrients will slowly diffuse away from the source into the surrounding water
(intestinal fluid) in such a way as to produce a diffusion gradient (or slope in concentration) with the
concentration increasing closer to the source and eventually dwindling to zero far from the source. Such
gradients are generally what we call exponential gradients, like the ones shown below:
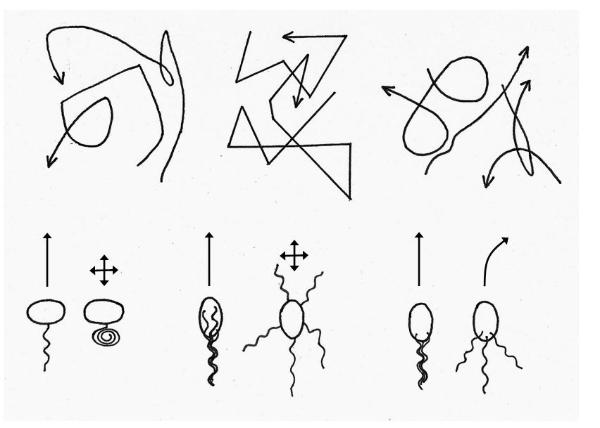
The diagram below compares the strategy used by Escherichia coli (middle) to that used by the
bacterium Rhodobacter sphaeroides (left) and Sinorhizobium meliloti (right). The top panel shows
typical traces of the movements of individual bacteria, whilst the bottom panel shows a pair of cells
with the left-most cell swimming forward and the right-most turning.
bacterium Rhodobacter sphaeroides (left) and Sinorhizobium meliloti (right). The top panel shows
typical traces of the movements of individual bacteria, whilst the bottom panel shows a pair of cells
with the left-most cell swimming forward and the right-most turning.
Redrawn from Armitage and Schmitt, 1997. Left: Rhodobacter sphaeroides, middle:
Escherichia coli and right: Sinorhizobium meliloti.
Escherichia coli and right: Sinorhizobium meliloti.
Rhodobacter sphaeroides has a unidirectional motor (it rotates in one direction only) which can change
speed and stops periodically. When the motor stops, the flagellum relaxes into a coil which slowly
rotates, turning the cell. Sinorhizobium meliloti has a bundle of flagella that only rotate CW but the
speed of rotation of the motors can change, causing the filaments to separate and the cell to turn
smoothly. The flagella never stop. Note that Escherichia coli turns abruptly by random amounts at
intervals, and so traces out a zig-zag path.
Modelling Chemokinesis
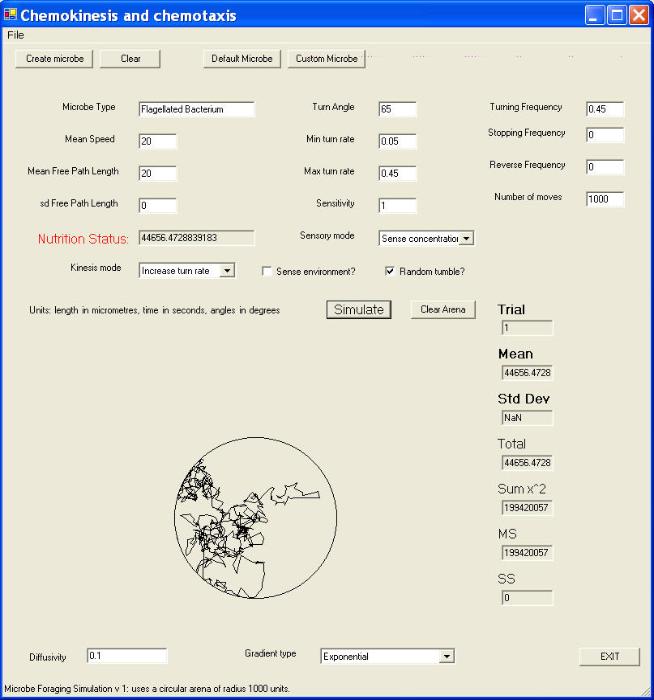
We shall now take a closer look at the strategy employed by Escherichia coli in particular. To assist in
this I shall use results obtained from a computer simulation, using a program that I wrote (in Visual C#) a
screenshot of which is shown below (click image to enlarge):
speed and stops periodically. When the motor stops, the flagellum relaxes into a coil which slowly
rotates, turning the cell. Sinorhizobium meliloti has a bundle of flagella that only rotate CW but the
speed of rotation of the motors can change, causing the filaments to separate and the cell to turn
smoothly. The flagella never stop. Note that Escherichia coli turns abruptly by random amounts at
intervals, and so traces out a zig-zag path.
Modelling Chemokinesis
We shall now take a closer look at the strategy employed by Escherichia coli in particular. To assist in
this I shall use results obtained from a computer simulation, using a program that I wrote (in Visual C#) a
screenshot of which is shown below (click image to enlarge):
See also: How do bacteria sense their environment?
To understand the different ways in which bacteria employ their flagella, we need to look at some of the
different patterns of flagellation as shown in the diagram below. In monotrichous flagellation, the
bacterium has a single flagellum, which may or may not be inserted at one of the narrow ends or poles
of the cell. In lophotrichous falgellation, a bundle of flagella is inserted into one of the cell poles. In
amphitrichous flagellation, one or more flagella are inserted at both poles. Finally, in peritrichous
flagellation the flagella are inserted at various places around the cell, apparently at random. These
arrangements depend upon the species or strain (subspecies) of the bacterium.
To understand the different ways in which bacteria employ their flagella, we need to look at some of the
different patterns of flagellation as shown in the diagram below. In monotrichous flagellation, the
bacterium has a single flagellum, which may or may not be inserted at one of the narrow ends or poles
of the cell. In lophotrichous falgellation, a bundle of flagella is inserted into one of the cell poles. In
amphitrichous flagellation, one or more flagella are inserted at both poles. Finally, in peritrichous
flagellation the flagella are inserted at various places around the cell, apparently at random. These
arrangements depend upon the species or strain (subspecies) of the bacterium.
I don't intend to explain all the features of this program (which is still under development) but only some
of the things it can do. Note that we are simulating the movements of bacteria in a circular arena (one
millimetre in diameter in the example above). The computer assigns a set-point value of the turning
frequency of 0.45 (which can be set by the user). The set-point is the default probability of turning
(which gives us the turning frequency) in the absence of any stimulus. This is the probability each
second that the bacterium will turn, in the case of Escherichia coli (modelled here) the probability each
second that it will switch its flagella into CCW rotation and tumble and so change direction to a new
random direction. The simulation here is two-dimensional, but extending to three dimensions should not
alter the physical principles (if we used a spherical arena in place of a circular one). Alternatively we can
make the microbe always turn by a fixed angle, such as 65 degrees. We can start the microbe from any
position in the arena, in this case from the centre. We can place a nutrient source in the arena, again in
the centre in this case. We can then tell the bacterium how to respond to the nutrient. First of all, let's
look at some examples when the bacteria ignores the nutrient (or equivalently if their is no nutrient
source in the arena).
of the things it can do. Note that we are simulating the movements of bacteria in a circular arena (one
millimetre in diameter in the example above). The computer assigns a set-point value of the turning
frequency of 0.45 (which can be set by the user). The set-point is the default probability of turning
(which gives us the turning frequency) in the absence of any stimulus. This is the probability each
second that the bacterium will turn, in the case of Escherichia coli (modelled here) the probability each
second that it will switch its flagella into CCW rotation and tumble and so change direction to a new
random direction. The simulation here is two-dimensional, but extending to three dimensions should not
alter the physical principles (if we used a spherical arena in place of a circular one). Alternatively we can
make the microbe always turn by a fixed angle, such as 65 degrees. We can start the microbe from any
position in the arena, in this case from the centre. We can place a nutrient source in the arena, again in
the centre in this case. We can then tell the bacterium how to respond to the nutrient. First of all, let's
look at some examples when the bacteria ignores the nutrient (or equivalently if their is no nutrient
source in the arena).
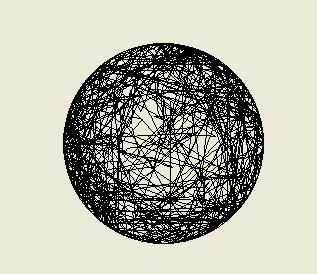
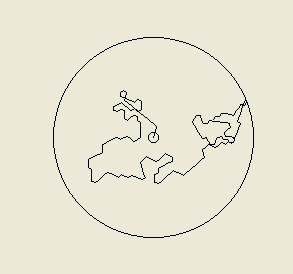
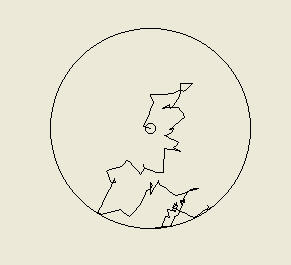
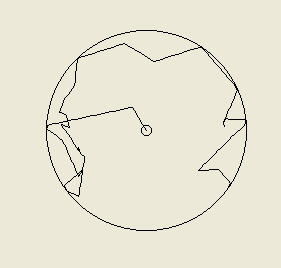
Above: the tracks of two computer simulated bacteria. On the left we have a bacterium that is making
fixed 65 degree turns (left or right) with a probability of 0.45 (45%) each second and moving at 20
micrometres per second. The path will be different every time the simulation is run, since every second
the computer determines whether or not the bacteria turns by 'rolling dice' as it were (it is a stochastic or
probabilistic model). On the right is a more realistic trace of what a cell of Escherichia coli might do, as
this bacterium is programmed to make tumbles and so turns by a random amount rather than a fixed
angle. This is what we call a random walk. A particle of smoke or a molecule of oxygen in still air would
move in a similar fashion, turning as gas molecules collide with it. If random walking was all these
bacteria did then moving about would be of limited use, they would disperse alright, but they would not
move toward favourable conditions, like a nice source of aspartate, they would effectively drift at
random. In the absence of any clear stimulus this is exactly what they do, but they have sensors always
on the alert. These sensors are called receptors, since they bind to nutrients, such as aspartate and
then alert the cell that a molecule of aspartate has been detected.
The question is, how do the bacteria go from detecting a nutrient to locating its source?
If the bacterium was fairly large, like an amoeba, which may be 100 micrometres (0.1 millimetres) across
or more, then it could sample the diffusion gradient spatially as shown in the two diagrams below (click
each figure to enlarge it):
fixed 65 degree turns (left or right) with a probability of 0.45 (45%) each second and moving at 20
micrometres per second. The path will be different every time the simulation is run, since every second
the computer determines whether or not the bacteria turns by 'rolling dice' as it were (it is a stochastic or
probabilistic model). On the right is a more realistic trace of what a cell of Escherichia coli might do, as
this bacterium is programmed to make tumbles and so turns by a random amount rather than a fixed
angle. This is what we call a random walk. A particle of smoke or a molecule of oxygen in still air would
move in a similar fashion, turning as gas molecules collide with it. If random walking was all these
bacteria did then moving about would be of limited use, they would disperse alright, but they would not
move toward favourable conditions, like a nice source of aspartate, they would effectively drift at
random. In the absence of any clear stimulus this is exactly what they do, but they have sensors always
on the alert. These sensors are called receptors, since they bind to nutrients, such as aspartate and
then alert the cell that a molecule of aspartate has been detected.
The question is, how do the bacteria go from detecting a nutrient to locating its source?
If the bacterium was fairly large, like an amoeba, which may be 100 micrometres (0.1 millimetres) across
or more, then it could sample the diffusion gradient spatially as shown in the two diagrams below (click
each figure to enlarge it):
Click figure to enlarge.
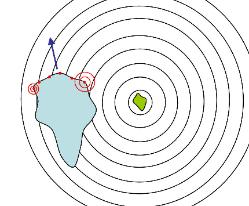
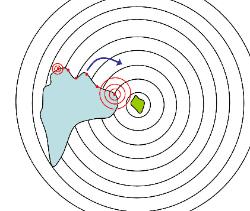
Above: left - a large cell, such as an amoeba or human white blood cell, has detected food (such as
bacteria, shown in green in the centre of the circles) releasing a gradient of chemicals (shown as
circular waves) whilst it was crawling along in the direction shown by the blue arrow. As the chemical
diffuses from the source its concentration drops (exponentially) and so receptors (the red dots at the
front of the cell) on the right of the cell detect a stronger signal than receptors on the left. This enables
the cell to compute that the source is located off its right and so the cell can turn and move more or less
straight toward the source. This movement directly toward a chemical source is called positive
chemotaxis (negative chemotaxis is movement directly away from a chemical source, such as a toxin or
irritant).
Most bacteria are much smaller than an animal cell or an amoeba, however, with a typical length of only
two micrometres and a diameter of one micrometre - one hundred times the diameter of our amoeba.
This makes it hard for a bacterium to detect differences in stimulus strength between its left and right
sides as the chemical has not diffused much to cross the width of the cell. Thus, it is thought that
bacteria do not use this method.
Another way to arrive at the source, if one cannot detect where it is coming from, is to turn about at
random (a random walk) but alter the probability of turning (or the turning frequency) according to the
concentration of the nutrient detected at each point in space (a mathematical point has no diameter)
and so undergo a biased walk. We know that Escherichia coli reduces its tumbling frequency in the
presence of nutrients, so let's add this to a model and see what happens, let's make our simulated
bacterium turn less frequently when it is in a region of high nutrient concentration. The results are
shown below:
bacteria, shown in green in the centre of the circles) releasing a gradient of chemicals (shown as
circular waves) whilst it was crawling along in the direction shown by the blue arrow. As the chemical
diffuses from the source its concentration drops (exponentially) and so receptors (the red dots at the
front of the cell) on the right of the cell detect a stronger signal than receptors on the left. This enables
the cell to compute that the source is located off its right and so the cell can turn and move more or less
straight toward the source. This movement directly toward a chemical source is called positive
chemotaxis (negative chemotaxis is movement directly away from a chemical source, such as a toxin or
irritant).
Most bacteria are much smaller than an animal cell or an amoeba, however, with a typical length of only
two micrometres and a diameter of one micrometre - one hundred times the diameter of our amoeba.
This makes it hard for a bacterium to detect differences in stimulus strength between its left and right
sides as the chemical has not diffused much to cross the width of the cell. Thus, it is thought that
bacteria do not use this method.
Another way to arrive at the source, if one cannot detect where it is coming from, is to turn about at
random (a random walk) but alter the probability of turning (or the turning frequency) according to the
concentration of the nutrient detected at each point in space (a mathematical point has no diameter)
and so undergo a biased walk. We know that Escherichia coli reduces its tumbling frequency in the
presence of nutrients, so let's add this to a model and see what happens, let's make our simulated
bacterium turn less frequently when it is in a region of high nutrient concentration. The results are
shown below:
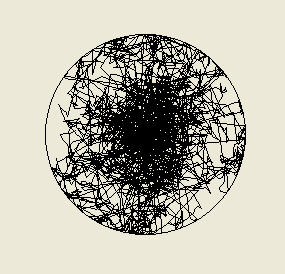
Above: the traces of ten bacteria simulated over 1000 seconds with each cell moving at a fixed speed of
20 micrometres per second. The nutrient source is diffusing from the centre of the arena and the bacteria
start at the centre and reduce their probability of turning in regions of high nutrient concentration (below
the set point probability of 0.45 per second) as does Escherichia coli, but whereas Escherichia coli is
attracted to nutrients, our model bacterium has been repelled - the bacteria accumulate far away from the
source where the nutrient concentration is very low, which is not very good if you want to absorb these
nutrients!
So what went wrong with our model? Apart from measuring concentration at various points in space there
is something else that a bacterium may be able to do - it may be able to compute a temporal comparison.
The bacterium could remember what the concentration was, say one second earlier, and then compare
that value with the current concentration in its new position (20 micrometres from where it was one second
ago) and determine whether the concentration has gone up or down. If the concentration has gone up
then it is now nearer to the source and so it would reduce its turning frequency. Let's give our bacteria
each a one second memory and see what happens:
20 micrometres per second. The nutrient source is diffusing from the centre of the arena and the bacteria
start at the centre and reduce their probability of turning in regions of high nutrient concentration (below
the set point probability of 0.45 per second) as does Escherichia coli, but whereas Escherichia coli is
attracted to nutrients, our model bacterium has been repelled - the bacteria accumulate far away from the
source where the nutrient concentration is very low, which is not very good if you want to absorb these
nutrients!
So what went wrong with our model? Apart from measuring concentration at various points in space there
is something else that a bacterium may be able to do - it may be able to compute a temporal comparison.
The bacterium could remember what the concentration was, say one second earlier, and then compare
that value with the current concentration in its new position (20 micrometres from where it was one second
ago) and determine whether the concentration has gone up or down. If the concentration has gone up
then it is now nearer to the source and so it would reduce its turning frequency. Let's give our bacteria
each a one second memory and see what happens:
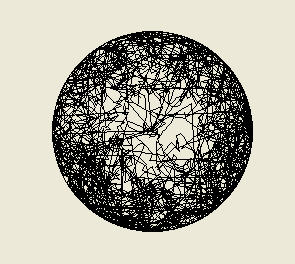
Above: the traces of ten simulated bacteria over 1000 seconds, in which each bacterium has a one
second memory, allowing it to compare concentrations of nutrient at two different positions and times
and so determine if it is getting nearer or further away. Now, by reducing their tumbling frequency as
they get nearer to the food in the centre (and increasing it up to the maximum set-point value of
0.45/second should they move away) the bacteria have found the source and spend most of their time
there. They still wander off a bit, which is good, because they may find another stronger source nearby!
Just to convince you further, look at the traces below:
second memory, allowing it to compare concentrations of nutrient at two different positions and times
and so determine if it is getting nearer or further away. Now, by reducing their tumbling frequency as
they get nearer to the food in the centre (and increasing it up to the maximum set-point value of
0.45/second should they move away) the bacteria have found the source and spend most of their time
there. They still wander off a bit, which is good, because they may find another stronger source nearby!
Just to convince you further, look at the traces below:
Above: left - the cells have been programmed to increase their turning frequency as they approach
the source, which causes them to avoid it. Right - with no source present, the cells move around the
whole arena at random and cover it more or less evenly.
Typical traces of single cells (over 200 seconds) are shown below:
the source, which causes them to avoid it. Right - with no source present, the cells move around the
whole arena at random and cover it more or less evenly.
Typical traces of single cells (over 200 seconds) are shown below:
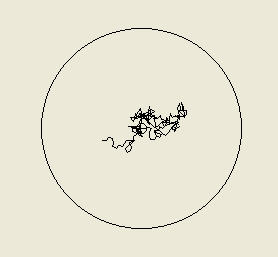
Above: left - a cell programmed to increase its turning frequency as it moves toward the source (and to
turn less as it moves away) avoids the source (it began where the source is located in the centre circle) as
it tends to turn away when it moves toward the source (up the gradient). Right - a bacterium programmed
to turn less often as it nears the source, and to turn more often as it moves away (up to the maximum
average of 0.45 turns / second) tends to turn back when if it moves away from the source and so is
attracted to the source. Remember that the bacteria here are responding to the direction they are moving
up or down the gradient rather than to the instantaneous nutrient concentration.
Thus, if we give the bacteria a one second memory then, in this situation, they are able to locate the
nutrients successfully! Other modellers have come to the same conclusion - that the bacteria probably
have a memory of up to several seconds. This way of locating nutrients by altering velocity (speed or
bearing) is called chemokinesis.
These simulations are ongoing and more needs to be done to make the simulations more realistic.
Eventually the plan is to use the simulator to model directed movement in other kinds of organisms,
including insects.
A note on terminology - chemotaxis or chemokinesis?
The traditional definitions of these terms, which are still used in most biological disciplines, such as
zoology, botany and animal behaviourial studies (ethology) are given in many standard textbooks as:
Chemotaxis: movement directly toward or away from a chemical source (accomplished by the
simultaneous sampling of gradients at more than one position in space, that is by measuring spatial
gradients directly).
Chemokinesis: an alteration in velocity (turning frequency or speed) in response to a chemical gradient.
This occurs when an organism moving through the gradient compares the stimulus strength at different
positions in space at different times, and so measures the gradient indirectly. This requires some form of
memory.
Organisms large enough to simultaneously compare stimulus strength at two different positions in space
can undergo taxis. Organisms too small to do this must undergo kinesis.
Thus, in bacteria we have the process of chemokinesis. In larger animal cells we actually have both
chemotaxis and chemokinesis. However, the vast majority of the microbiology literature refers to
chemotaxis in bacteria and the two terms have been massively confused, making the terms almost
useless! After all, is sampling over time really so different to sampling in space when the end result is the
same? Also, since taxis involves a change in velocity as the organism turns toward the source it is
arguably a form of chemokinesis . 'Kinesis' simply refers to motion, in which case chemotaxis is a form of
chemokinesis! What I am saying is that the terms are unfortunate, since 'kinetic' simply refers to motion of
any kind.
Chemotaxis does not seem the correct term to apply to most bacteria, in my opinion, and I shall refer to
chemokinesis in the case of bacteria (contrary to the mainstream scientific literature).
Chemotaxis is the direct movement towards or away from a chemical source, for example in amoebae,
white blood cells and insects, which are large enough to sample stimulus concentration at different
positions in space simultaneously and hence determine the direction of the source.
Chemokinesis is not precise movement towards or away from a chemical source, but involves reaching a
destination without being truly orientated. All that is required are sense organs that determine
concentration at a single position in space at any one time. Combined with memory (as we saw above) or
habituation, this allows an organism to respond to a stimulus concentration gradient, by detecting changes
over time as the organism moves, even though the immediate response is undirected. In orthokinesis,
the average speed of movement or the frequency of activity depends on the intensity of stimulation. In
klinokinesis the organism turns in a straight line in a favourable environment, but as soon as the
environment becomes less favourable it begins making turns, until, after a time, the organism adapts or
habituates and begins moving in a straight line again (this is a form of memory that allows the organism to
detect changes in stimulus intensity, i.e. gradients). Random turning and adaptation eventually delivers the
organism to the favourable environment, but by a stochastic process not a strictly directional one. What we
see in Escherichia coli is klinokinesis, though some bacteria also exhibit orthokinesis.
The modern usage of the terms in bacteriology is at odds with the classical biological textbook definitions
as stated above. Chemotaxis is taken to mean turning in response to a stimulus (in the sense of
klinokinesis) whilst chemokinesis is taken to mean a change in speed (orthokinesis). No distinction is then
made with more directional responses, such as an insect moving directly toward a source (chemotaxis). It
is easy to see how such confusion came about and this is unfortunate. On this website we prefer
consistency across the sub-disciplines of biology and so stick to the classical definitions. I can, however,
see why these somewhat clumsy terms have become confused! The term 'kinetic' simply refers to motion,
and so one has to wonder whether it made sense to set-up the term taxis and kinesis as opposing
opposites, when taxis still involves motion! The key is to understand that organisms can locate a stimulus
source either by directly measuring spatial gradients (in stereo) as humans can when they perceive a
sound, by comparing the stimulus intensity or loudness at each ear, or they can instead measure changes
in stimulus intensity over time and indirectly measure spatial gradients by comparing how the stimulus
varies over time as they change position. Almost all bacteria must take the latter approach, since they are
too small to measure gradients directly 9with few exceptions).
We can then recognise several types of movement in response to a chemical (terminology aside):
1. That due to spatial sampling and so rather directed straight toward or away from the source.
2. That due to a change in turning frequency and/or speed due to the instantaneous chemical
concentration.
3. That due to temporal sampling of the chemical gradient resulting in movement up or down a chemical
gradient by changing turning frequency and/or speed.
4. A combination of the above.
Is it only chemicals that attract or repel bacteria?
No, bacteria may respond to a wide variety of stimuli, including touch, light, temperature and magnetic
fields (e.g. toward or away from the magnetic North Pole) - different species respond differently according
to their needs and not all species respond to the same range of stimuli. Also, chemicals that attract
bacteria may not simply be nutrients, some bacteria are attracted by oxygen, whilst those that find oxygen
poisonous are possibly repelled by it . Some bacteria require sulphur or other chemicals in place of oxygen
and may be attracted toward these.
Bacteria may also show other responses to stimuli, apart from kinetic responses - they may secrete
materials, switch genes on or off and so synthesise new proteins, etc. To perform appropriate reactions in
complex environments bacteria must perform computations on all the incoming sensory data they receive -
each bacterium, minute as it is, is a very sophisticated computer!
In future updates we shall look at the sensors of bacteria in more detail.
Back to bacteria introduction...
Back to bacterial motility...
turn less as it moves away) avoids the source (it began where the source is located in the centre circle) as
it tends to turn away when it moves toward the source (up the gradient). Right - a bacterium programmed
to turn less often as it nears the source, and to turn more often as it moves away (up to the maximum
average of 0.45 turns / second) tends to turn back when if it moves away from the source and so is
attracted to the source. Remember that the bacteria here are responding to the direction they are moving
up or down the gradient rather than to the instantaneous nutrient concentration.
Thus, if we give the bacteria a one second memory then, in this situation, they are able to locate the
nutrients successfully! Other modellers have come to the same conclusion - that the bacteria probably
have a memory of up to several seconds. This way of locating nutrients by altering velocity (speed or
bearing) is called chemokinesis.
These simulations are ongoing and more needs to be done to make the simulations more realistic.
Eventually the plan is to use the simulator to model directed movement in other kinds of organisms,
including insects.
A note on terminology - chemotaxis or chemokinesis?
The traditional definitions of these terms, which are still used in most biological disciplines, such as
zoology, botany and animal behaviourial studies (ethology) are given in many standard textbooks as:
Chemotaxis: movement directly toward or away from a chemical source (accomplished by the
simultaneous sampling of gradients at more than one position in space, that is by measuring spatial
gradients directly).
Chemokinesis: an alteration in velocity (turning frequency or speed) in response to a chemical gradient.
This occurs when an organism moving through the gradient compares the stimulus strength at different
positions in space at different times, and so measures the gradient indirectly. This requires some form of
memory.
Organisms large enough to simultaneously compare stimulus strength at two different positions in space
can undergo taxis. Organisms too small to do this must undergo kinesis.
Thus, in bacteria we have the process of chemokinesis. In larger animal cells we actually have both
chemotaxis and chemokinesis. However, the vast majority of the microbiology literature refers to
chemotaxis in bacteria and the two terms have been massively confused, making the terms almost
useless! After all, is sampling over time really so different to sampling in space when the end result is the
same? Also, since taxis involves a change in velocity as the organism turns toward the source it is
arguably a form of chemokinesis . 'Kinesis' simply refers to motion, in which case chemotaxis is a form of
chemokinesis! What I am saying is that the terms are unfortunate, since 'kinetic' simply refers to motion of
any kind.
Chemotaxis does not seem the correct term to apply to most bacteria, in my opinion, and I shall refer to
chemokinesis in the case of bacteria (contrary to the mainstream scientific literature).
Chemotaxis is the direct movement towards or away from a chemical source, for example in amoebae,
white blood cells and insects, which are large enough to sample stimulus concentration at different
positions in space simultaneously and hence determine the direction of the source.
Chemokinesis is not precise movement towards or away from a chemical source, but involves reaching a
destination without being truly orientated. All that is required are sense organs that determine
concentration at a single position in space at any one time. Combined with memory (as we saw above) or
habituation, this allows an organism to respond to a stimulus concentration gradient, by detecting changes
over time as the organism moves, even though the immediate response is undirected. In orthokinesis,
the average speed of movement or the frequency of activity depends on the intensity of stimulation. In
klinokinesis the organism turns in a straight line in a favourable environment, but as soon as the
environment becomes less favourable it begins making turns, until, after a time, the organism adapts or
habituates and begins moving in a straight line again (this is a form of memory that allows the organism to
detect changes in stimulus intensity, i.e. gradients). Random turning and adaptation eventually delivers the
organism to the favourable environment, but by a stochastic process not a strictly directional one. What we
see in Escherichia coli is klinokinesis, though some bacteria also exhibit orthokinesis.
The modern usage of the terms in bacteriology is at odds with the classical biological textbook definitions
as stated above. Chemotaxis is taken to mean turning in response to a stimulus (in the sense of
klinokinesis) whilst chemokinesis is taken to mean a change in speed (orthokinesis). No distinction is then
made with more directional responses, such as an insect moving directly toward a source (chemotaxis). It
is easy to see how such confusion came about and this is unfortunate. On this website we prefer
consistency across the sub-disciplines of biology and so stick to the classical definitions. I can, however,
see why these somewhat clumsy terms have become confused! The term 'kinetic' simply refers to motion,
and so one has to wonder whether it made sense to set-up the term taxis and kinesis as opposing
opposites, when taxis still involves motion! The key is to understand that organisms can locate a stimulus
source either by directly measuring spatial gradients (in stereo) as humans can when they perceive a
sound, by comparing the stimulus intensity or loudness at each ear, or they can instead measure changes
in stimulus intensity over time and indirectly measure spatial gradients by comparing how the stimulus
varies over time as they change position. Almost all bacteria must take the latter approach, since they are
too small to measure gradients directly 9with few exceptions).
We can then recognise several types of movement in response to a chemical (terminology aside):
1. That due to spatial sampling and so rather directed straight toward or away from the source.
2. That due to a change in turning frequency and/or speed due to the instantaneous chemical
concentration.
3. That due to temporal sampling of the chemical gradient resulting in movement up or down a chemical
gradient by changing turning frequency and/or speed.
4. A combination of the above.
Is it only chemicals that attract or repel bacteria?
No, bacteria may respond to a wide variety of stimuli, including touch, light, temperature and magnetic
fields (e.g. toward or away from the magnetic North Pole) - different species respond differently according
to their needs and not all species respond to the same range of stimuli. Also, chemicals that attract
bacteria may not simply be nutrients, some bacteria are attracted by oxygen, whilst those that find oxygen
poisonous are possibly repelled by it . Some bacteria require sulphur or other chemicals in place of oxygen
and may be attracted toward these.
Bacteria may also show other responses to stimuli, apart from kinetic responses - they may secrete
materials, switch genes on or off and so synthesise new proteins, etc. To perform appropriate reactions in
complex environments bacteria must perform computations on all the incoming sensory data they receive -
each bacterium, minute as it is, is a very sophisticated computer!
In future updates we shall look at the sensors of bacteria in more detail.
Back to bacteria introduction...
Back to bacterial motility...
Above: monotrichous flagellation (one flagella) which may be polar, as on the left (inserted into one of
the cell poles) or non-polar as on the right.
the cell poles) or non-polar as on the right.
Above: lophotrichous flagellation: several flagella are inserted into one cell pole.
Above: amphitrichous flagellation: one or more flagella are inserted into each pole (also called bipolar
flagellation).
flagellation).
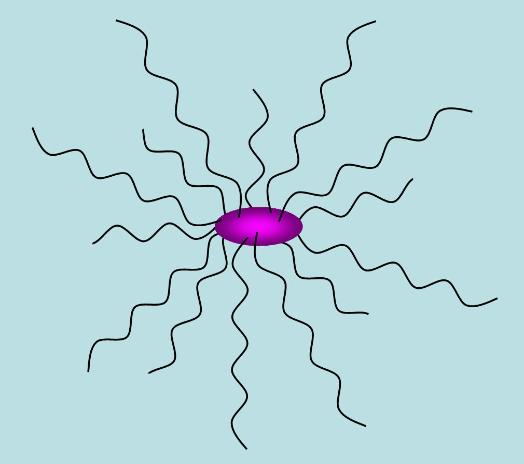
Above: peritrichous flagellation: several flagella emerge from random positions on the cell surface, but
come together as a bundle at one end of the cell in order to propel it.
come together as a bundle at one end of the cell in order to propel it.
Above: peritrichous flagellation in which the flagella have flown apart from their bundle, causing the cell
to tumble and change orientation.
to tumble and change orientation.